Where does XbaI restriction enzyme cut?

Daniel Rodriguez
Published Mar 19, 2026
Where does XbaI restriction enzyme cut?
37°C
Thermo Scientific XbaI restriction enzyme recognizes T^CTAGA sites and cuts best at 37°C in Tango buffer. See Reaction Conditions for Restriction Enzymes for a table of enzyme activity, conditions for double digestion, and heat inactivation for this and other restriction enzymes.
Where does the restriction enzyme HindIII cut?
HindIII (pronounced “Hin D Three”) is a type II site-specific deoxyribonuclease restriction enzyme isolated from Haemophilus influenzae that cleaves the DNA palindromic sequence AAGCTT in the presence of the cofactor Mg2+ via hydrolysis.
What will cut restriction enzymes?
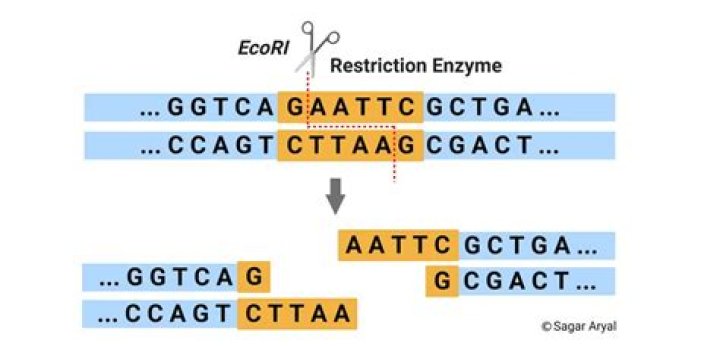
Restriction enzymes cut through both nucleotide strands, breaking the DNA into fragments, but they don’t always do this in the same way. This overhanging nucleotide strand is called a sticky end because it can easily bond with complementary DNA fragments.
What do Type 2 restriction enzymes cut?
Type II enzymes cut DNA at defined positions close to or within their recognition sequences. They produce discrete restriction fragments and distinct gel banding patterns, and they are the predominant class used in the laboratory for routine DNA analysis and gene cloning.
How does XbaI cut?
XbaI is blocked by overlapping methylation, directed by the GATC sequence (methylation of the adenine moiety), in E. If you propagate your plasmid in a methylation-deficient host such as JM110, you’re plasmid will cut normally with XbaI. In methylation-competent strains, no digestion will be possible.
Does XbaI produce sticky ends?
Note that both XbaI and SpeI have the same sticky ends, CTAG. As a result, DNA cut by one enzyme can stick to DNA cut by the other enzyme.
Does HindIII produce blunt ends?
Cutting DNA with restriction enzymes can produce fragments with either blunt or sticky ends. HindIII – recognises the sequence 5’AAGCTT’3 – sticky ends. PstI – recognises the sequence 5’CTGCAG’3 – sticky ends. Sau3A – recognises the sequence 5’GATC’3 (produces the same sticky ends as BamHI upon cutting)
What temperature does HindIII work best in and why?
Thermo Scientific HindIII restriction enzyme recognizes A^AGCTT sites and cuts best at 37°C in R buffer. See Reaction Conditions for Restriction Enzymes for a table of enzyme activity, conditions for double digestion, and heat inactivation for this and other restriction enzymes.
Why are restriction enzymes called molecular scissors?
Restriction enzymes are also called “molecular scissors” as they cleave DNA at or near specific recognition sequences known as restriction sites. These enzymes make one incision on each of the two strands of DNA and are also called restriction endonucleases.
Why would a restriction enzyme not cut?
The preparation of DNA to be cleaved should be free of contaminants such as phenol, chloroform, alcohol, EDTA, detergents, or excessive salts, all of which can interfere with restriction enzyme activity. If an inhibitor (often salt, EDTA or phenol) is present, the control DNA will not cut after mixing.
What is Type 4 restriction enzyme?
As for their recognition pattern, type IV enzymes usually recognize asymmetrical DNA sequences and ENase cleavage occurs at a defined distance from the recognition site, as in type IIS enzymes. An exception is R. HaeIV, where cleavage occurs on both sides of the recognition site.