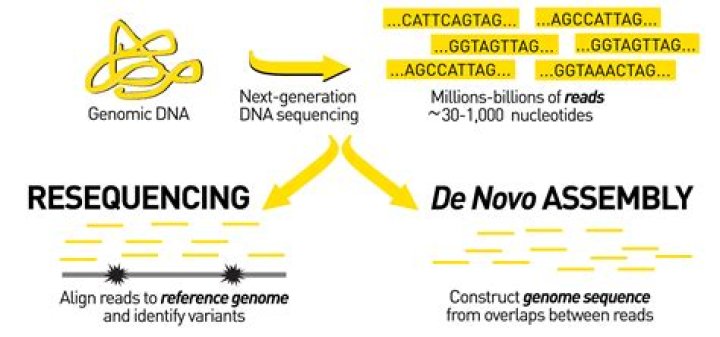
What is de novo assembly of sequences

John Castro
Published Apr 28, 2026
De novo sequencing refers to sequencing a novel genome where there is no reference sequence available for alignment. Sequence reads are assembled as contigs, and the coverage quality of de novo sequence data depends on the size and continuity of the contigs (ie, the number of gaps in the data).
What is de novo gene assembly?
De novo genome assembly is a strategy for genome assembly, representing the genome assembly of a novel genome from scratch without the aid of reference genomic data. De novo genome assemblies assume no prior knowledge of the source DNA sequence length, layout or composition.
What is contig in bioinformatics?
A contig–from the word “contiguous”–is a series of overlapping DNA sequences used to make a physical map that reconstructs the original DNA sequence of a chromosome or a region of a chromosome. A contig can also refer to one of the DNA sequences used in making such a map.
What is de novo assembly used for?
De novo sequence assemblers are a type of program that assembles short nucleotide sequences into longer ones without the use of a reference genome. These are most commonly used in bioinformatic studies to assemble genomes or transcriptomes.What is de novo analysis?
Introduction. De novo (from new) genome assembly refers to the process of reconstructing an organism’s genome from smaller sequenced fragments. … Coverage can simply be computed by dividing the total number of sequenced bases by the “expected” size of the genome in question.
What do you mean by sequencing?
the process of combining things in a particular order, or discovering the order in which they are combined: A common sign of dyslexia is that the sequencing of letters when spelling words may be incorrect.
How does de novo sequencing work?
De novo is Latin which means “over again” or “anew”. The de novo peptide sequencing is a method for peptide sequencing performed without prior knowledge of the amino acid sequence. … In this method, the peptide is fragmented along the peptide backbone and the resulting fragment ions are measured to produce spectra.
What is genome sequencing?
Genome sequencing is figuring out the order of DNA nucleotides, or bases, in a genome—the order of As, Cs, Gs, and Ts that make up an organism’s DNA. … Much as your eye scans a sequence of letters to read a sentence, these machines “read” a sequence of DNA bases.Why is genome assembly important?
Assembly is required, because sequence read lengths – at least for now – are much shorter than most genomes or even most genes. … Although bacterial genomes are much smaller, genes are not necessarily in the same location and multiple copies of the same gene may appear in different locations on the genome.
What is Assembly in bioinformatics?In bioinformatics, sequence assembly refers to aligning and merging fragments from a longer DNA sequence in order to reconstruct the original sequence. … Typically the short fragments, called reads, result from shotgun sequencing genomic DNA, or gene transcript (ESTs).
Article first time published onWhat is N50 length?
Given a set of contigs, the N50 is defined as the sequence length of the shortest contig at 50% of the total genome length. … N50 can be described as a weighted median statistic such that 50% of the entire assembly is contained in contigs or scaffolds equal to or larger than this value.
How does contig assembly work?
The assembly of a contig map involves several steps. First, DNA is sheared into larger (50–200kb) pieces, which are cloned into BACs or PACs to form a BAC library. Since these clones should cover the entire genome/chromosome, it is theoretically possible to assemble a contig of BACs that covers the entire chromosome.
What is contig and scaffold?
A scaffold is a portion of the genome sequence reconstructed from end-sequenced whole-genome shotgun clones. … A contig is a contiguous length of genomic sequence in which the order of bases is known to a high confidence level.
What is de novo RNA synthesis?
De novo synthesis is a unique phase in the transcription cycle where the RNA polymerase binds two nucleotides rather than a nascent RNA polymer and a single nucleotide. … The structures show that the initiating nucleotides bind RNA polymerase in locations distinct from those described previously for elongation complexes.
What is metagenomic data?
Metagenomics is defined as the direct genetic analysis of genomes contained with an environmental sample. The field initially started with the cloning of environmental DNA, followed by functional expression screening [1], and was then quickly complemented by direct random shotgun sequencing of environmental DNA [2,3].
Which of these steps can be taken to improve genome assemblies?
- Investigate the properties of the genome you study. Every assembly or annotation project is different. …
- Extract high quality DNA. …
- Choose an appropriate sequencing technology. …
- Estimate the necessary computational resources. …
- Assemble your genome.
How do peptides fragment?
ETD fragments peptides by transferring en electron from radical anion to protonated peptide. This induces fragmentation along the backbone, generating c and z ions. The reaction allows cleavage without side chain chemistry, eg neutral loss.
What are peptides?
A peptide is a short chain of amino acids. The amino acids in a peptide are connected to one another in a sequence by bonds called peptide bonds. Typically, peptides are distinguished from proteins by their shorter length, although the cut-off number of amino acids for defining a peptide and protein can be arbitrary.
What are the 4 types of sequence?
- Arithmetic sequence.
- Geometric sequence.
- Harmonic sequence.
- Fibonacci sequence.
What are the 2 types of sequence?
- Arithmetic Sequences.
- Geometric Sequences.
- Harmonic Sequences.
- Fibonacci Numbers.
What does sequencing mean in computing?
What is sequencing? An explanation of sequencing, as used in algorithms and programming. Transcript. Algorithms consist of instructions that are carried out (performed) one after another. Sequencing is the specific order in which instructions are performed in an algorithm.
What is genome sequencing Covid?
Genome sequencing for COVID-19 is about developing a complete picture of a virus’s RNA. It involves obtaining positive COVID-19 samples and generating a complete RNA sequence of that virus from that sample.
What is genome assembly problem?
The basic problem of genome assembly stems from the fact that while genomes themselves are quite large and contain long stretches of contiguous sequence, on the order of millions of base pairs), the current generation of commonly used genome sequencers can only generate relatively short segments of sequence.
What is DNA assembly?
For the purposes of cloning, DNA assembly refers to a method to physically link together multiple fragments of DNA, in an end-to-end fashion, to achieve a desired, higher-order assembly prior to joining to a vector.
What genome means?
A genome is the complete set of genetic information in an organism. It provides all of the information the organism requires to function. In living organisms, the genome is stored in long molecules of DNA called chromosomes.
How genome sequencing is done?
Sequencing employs a technique known as electrophoresis to separate pieces of DNA that differ in length by only one base. In electrophoresis, DNA to be sequenced is placed at one end of a gel—a slab of a gelatin-like substance. (A major part of DNA sequencing simply comes down to making a bunch of Jell-O.)
Why is whole genome sequencing important?
It is ideal for discovery applications, such as identifying causative variants and novel genome assembly. Whole-genome sequencing can detect single nucleotide variants, insertions/deletions, copy number changes, and large structural variants.
What is assembly sequence planning?
Assembly sequence planning (ASP) is the process of computing a sequence of assembly motions for constituent parts of an assembled final product. ASP is proven to be NP-hard and thus its effective and efficient solution has been a challenge for the researchers in the field.
How is a DNA sequence assembled?
Assembly involves taking a large number of DNA reads, looking for areas in which they overlap with each other and then gradually piecing together the ‘jigsaw’. It is an attempt to reconstruct the original genome.
What does assembly sequence affect in the Assembly?
Assembly sequence planning (ASP) is the basis and critical step in the assembly process design. The selection of the assembly sequence has great effect on assembly feasibility, assembly complexity and assembly accuracy. Reasonable ASP can ensure assembly ability, reduce assembly costs, and shorten assembly time.
What is N50 value in sequencing?
N50 is the shortest contig length that needs to be included for covering 50% of the genome. Meaning. -> Half of the genome sequence is covered by contigs larger than or equal the N50 contig size. -> The sum of the lengths of all contigs of size N50 or longer contain at least 50 percent of the total genome sequence.



