What is an example of alternative splicing?

Michael Henderson
Published Mar 11, 2026
What is an example of alternative splicing?
Alternative splicing is a powerful means of controlling gene expression and increasing protein diversity. The best example is the Drosophila Down syndrome cell adhesion molecule (Dscam) gene, which can generate 38,016 isoforms by the alternative splicing of 95 variable exons.
Can Lncrna be spliced?
lncRNAs, unlike eRNAs, are usually spliced (De Santa et al., 2010; Ulitsky and Bartel, 2013), as are >90% of the lncRNAs associated to an EPC in our datasets.
Are Lncrna introns?
These retained introns occur in over 50% of lncRNAs examined and are mostly first introns with an average length of just 354 bp, compared to the average length of all human introns of 6355 and 7987 bp in mRNAs and lncRNAs, respectively.
Are non coding RNAs alternatively spliced?
AS (alternative splicing) is a fundamental process by which a gene can generate multiple distinct mRNA transcripts to increase protein diversity. NcRNAs (noncoding RNAs) are an abundant class of RNAs that do not encode proteins.
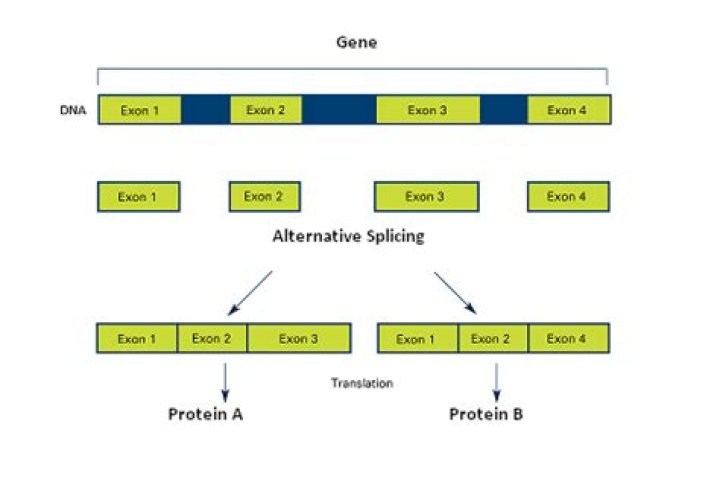
What is alternative splicing simple?
Alternative splicing is the process of selecting different combinations of splice sites within a messenger RNA precursor (pre-mRNA) to produce variably spliced mRNAs. These multiple mRNAs can encode proteins that vary in their sequence and activity, and yet arise from a single gene.
Is alternative splicing only in eukaryotes?
Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome; in humans, ~95% of multi-exonic genes are alternatively spliced.
Does lncRNA have ORF?
Many lncRNAs even have several shorter ORFs ranging in size from 100 nt to several hundred nucleotides within this region (Fig. Indeed, it has been shown that the presence of such upstream ORFs can lead to up to 80% reduction in translation efficiency and may even reduce the stability of the mRNAs (Calvo et al.
Are lncRNAs Polyadenylated?
Long non-coding RNAs (lncRNAs) are grouped into transcripts that are > 200 nucleotides in length. The human genome is estimated to contain ~16,000 lncRNA genes (Gencode 27)….Table 1.
| Type | Feature | Recommended reviews/articles |
|---|---|---|
| mRNA-like lncRNAs | 5′-capping and 3′ poly-A tails can be spliced | [149, 150] |
How do you identify lncRNA?
From here, novel lncRNAs can be identified by excluding protein-coding transcripts and annotated lncRNAs based on the databases of RefSeq, ENCODE, and FANTOM (Functional Annotation of the Mammalian Genome) [42], as well as the two databases of experimentally verified lncRNAs generated by the Mattick lab: lncRNAdb [43] …
Does lncRNA have Polya?
Long non-coding RNAs (lncRNAs) are grouped into transcripts that are > 200 nucleotides in length. The human genome is estimated to contain ~16,000 lncRNA genes (Gencode 27). Most of the lncRNAs contain normal 5′-caps and 3′ poly-A tails.
Does Lncrna have ORF?
How do you identify alternative splicing?
Quantification of alternative splicing to detect the abundance of differentially spliced isoforms of a gene in total RNA can be accomplished via RT-PCR using both quantitative real-time and semi-quantitative PCR methods.