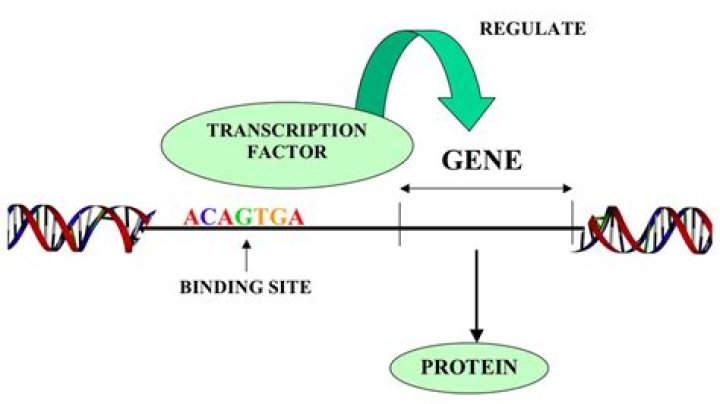
How genes are regulated transcription factors

Mia Smith
Published Apr 02, 2026
The activity of a transcription factor is often regulated by (de) phosphorylation, which may affect different functions, e.g. nuclear localization DNA binding and trans-activation. Ligand binding is another mode of transcription-factor activation. It is typical for the large super-family of nuclear hormone receptors.
How are transcription factors regulated?
The activity of a transcription factor is often regulated by (de) phosphorylation, which may affect different functions, e.g. nuclear localization DNA binding and trans-activation. Ligand binding is another mode of transcription-factor activation. It is typical for the large super-family of nuclear hormone receptors.
How are genes regulated in translation?
Gene expression is regulated at multiple levels, including the translation of mRNAs into proteins. … Most regulation is exerted at the first stage, where the AUG start codon is identified and decoded by the methionyl tRNA specialized for initiation (Met-tRNAi).
Are transcription factors transcriptional regulators?
The regulation of transcription is a vital process in all living organisms. It is orchestrated by transcription factors and other proteins working in concert to finely tune the amount of RNA being produced through a variety of mechanisms.How do transcription factors work what is their relationship to control regions in DNA?
Transcription factors are proteins possessing domains that bind to the DNA of promoter or enhancer regions of specific genes. They also possess a domain that interacts with RNA polymerase II or other transcription factors and consequently regulates the amount of messenger RNA (mRNA) produced by the gene.
Are transcription factors genes?
Transcription factor (TF) genes encode DNA-binding proteins. In all organisms, TFs play central roles in transcription by regulating gene expression. TFs are involved in a variety of biological processes, such as development and cell cycle control. TFs comprise one of the largest known groups of genes.
How is gene expression regulated after transcription?
The pre-mRNA has to go through some modifications to become a mature mRNA molecule that can leave the nucleus and be translated. These include splicing, capping, and addition of a poly-A tail, all of which can potentially be regulated – sped up, slowed down, or altered to result in a different product.
How is protein expression regulated?
Eukaryotic gene expression is regulated during transcription and RNA processing, which take place in the nucleus, and during protein translation, which takes place in the cytoplasm. Further regulation may occur through post-translational modifications of proteins.Why is regulation of transcription important?
Transcriptional regulation is a critical biological process that allows the cell or an organism to respond to a variety of intra- and extra-cellular signals, to define cell identity during development, to maintain it throughout its lifetime, and to coordinate cellular activity.
What are regulatory transcription factors quizlet?refers to the phenomenon that the level of gene expression can be controlled so that genes can be expressed at high or low levels. … Transcription factor. to describe proteins that influence the ability of RNA polymerase to transcribe a given gene.
Article first time published onHow are genes regulated?
Gene regulation can occur at any point during gene expression, but most commonly occurs at the level of transcription (when the information in a gene’s DNA is passed to mRNA). Signals from the environment or from other cells activate proteins called transcription factors.
How does attenuation regulate gene expression?
Like regulation by the trp repressor, attenuation is a mechanism for reducing expression of the trp operon when levels of tryptophan are high. However, rather than blocking initiation of transcription, attenuation prevents completion of transcription.
How is transcription regulated in eukaryotes?
Gene expression in eukaryotic cells is regulated by repressors as well as by transcriptional activators. Like their prokaryotic counterparts, eukaryotic repressors bind to specific DNA sequences and inhibit transcription. … Other repressors compete with activators for binding to specific regulatory sequences.
Do transcription factors control gene expression?
Transcription factors are proteins involved in the process of converting, or transcribing, DNA into RNA. … Regulation of transcription is the most common form of gene control. The action of transcription factors allows for unique expression of each gene in different cell types and during development.
How do transcription factors find their targets?
Transcription factors (which are described in the video) have to be able to first scan the genome so they can find their target sites and then bind there, which will turn genes on or off. … It’s known that they can also randomly attach to the genome non-specifically.
How do transcription factors bind DNA?
Response elements The DNA sequence that a transcription factor binds to is called a transcription factor-binding site or response element. Transcription factors interact with their binding sites using a combination of electrostatic (of which hydrogen bonds are a special case) and Van der Waals forces.
How do external factors change the Post transcription regulation of gene expression?
How can external stimuli alter post-transcriptional control of gene expression? External stimuli can modify RNA-binding proteins (i.e., through phosphorylation of proteins) to alter their activity.
How do operons regulate gene expression?
Genes in an operon are transcribed as a group and have a single promoter. Each operon contains regulatory DNA sequences, which act as binding sites for regulatory proteins that promote or inhibit transcription.
What are three mechanisms by which transcription factors regulate eukaryotic gene expression?
Sets of transcription factor proteins bind to specific DNA sequences in or near a gene and promote or repress its transcription into an RNA. RNA processing. Splicing, capping, and addition of a poly-A tail to an RNA molecule can be regulated, and so can exit from the nucleus.
How do transcription factors control protein synthesis at the transcription level?
Transcription factors (TFs) are regulatory proteins whose function is to activate (or more rarely, to inhibit) transcription of DNA by binding to specific DNA sequences. TFs have defined DNA-binding domains with up to 106-fold higher affinity for their target sequences than for the remainder of the DNA strand.
How do cells regulate genes using alternative splicing?
Alternative splicing can regulate protein composition by changing the coding content between isoforms of the same gene. As a consequence, AS contributes to increased protein diversity and, ultimately, cellular and functional complexity, without increasing the size of a eukaryotic organism’s genome (Stamm et al., 2005).
Are transcription factors enzymes?
The regulatory potential of such a bifunctional transcription factor is enormous and suggests a new layer of dynamic regulation in which transcription factors themselves might provide intrinsic enzymatic activities to fine-tune nuclear output.
In what two ways is gene regulation in eukaryotes different from gene regulation in prokaryotes?
Prokaryotic transcription and translation occur simultaneously in the cytoplasm, and regulation occurs at the transcriptional level. Eukaryotic gene expression is regulated during transcription and RNA processing, which take place in the nucleus, and during protein translation, which takes place in the cytoplasm.
How can you inhibit gene expression?
The genes can be silenced by siRNA molecules that cause the endonucleatic cleavage of the target mRNA molecules or by miRNA molecules that suppress translation of the mRNA molecule. With the cleavage or translational repression of the mRNA molecules, the genes that form them are rendered essentially inactive.
How can gene regulation conserve energy?
They can conserve energy and resources by regulating their activities, producing only those genes necessary for the cell to function. In prokaryotes, DNA-binding proteins regulate genes by controlling transcription. … Complex gene regulation in eukaryotes makes cell specialization possible.
What does epigenetic gene regulation rely on?
In Summary: Eukaryotic Epigenetic Gene Regulation Epigenetic mechanisms control access to the chromosomal region to allow genes to be turned on or off. These mechanisms control how DNA is packed into the nucleus by regulating how tightly the DNA is wound around histone proteins.
Which is the primary step for regulation of gene expression?
The regulation of transcription is the primary step for the regulation of gene expression.
How does genetic code affect gene expression?
Gene expression is the process the cell uses to produce the molecule it needs by reading the genetic code written in the DNA. To do this, the cell interprets the genetic code, and for each group of three letters it adds one of the 20 different amino acids that are the basic units needed to build proteins.
What is the difference between General transcription factors and regulatory transcription factors?
General transcription factors are a small group of highly abundant proteins that assemble on the promoters of all genes transcribed by RNA polymerase II, whereas thousands of different low abundant proteins are gene regulatory TFs.
What is a major difference between the General transcription factors and regulatory transcription factors?
What is a major difference between the general and regulatory transcription factors? General transcription factors are essential for any transcription for all genes while regulatory transcription factors regulate transcription of specific genes.
What's the difference between General transcription factors and regulatory transcription factors?
The General transcription factors are the factors which are used to form the pre-initiation complex during the process of transcription. The Specific transcription factors are either enhancers or repressors, which are specific DNA sequences that activate or repress the general transcription process.